Section 3 Point Cloud Processing
We’ll use the cloud2trees package to perform all preprocessing of point cloud data which includes:
- ground classification and noise removal
- raster data (DTM and CHM) generation
- point cloud height normalization
All of this can be accomplished using the cloud2trees::cloud2raster() function. After generating these products from the raw point cloud we’ll perform object segmentation to attempt to detect slash piles from the CHM which we’ll generate by setting the minimum height to zero (essentially a digital surface model [DSM] with the ground removed).
To attempt to identify slash piles from the CHM/DSM, we can use watershed segmentation or DBSCAN (Density-Based Spatial Clustering of Applications with Noise), for example. Both of these methods are common in segmenting individual trees and coarse woody debris from CHM data.
3.1 Process Raw Point Cloud
We’ll use cloud2trees::cloud2raster() to process the raw point cloud data. For each site we’ll generate a fine-resolution (0.1 m) CHM data set which can be aggregated to coarser resolution for sensitivity testing. Fine resolution CHM data would include granular details which may be important for delineating smaller slash piles while more coarse resolution data may help smooth out noise in the data to reduce false positive pile detections and potentially better represent the form of the actual piles
3.1.1 PSINF Mixed Conifer Site
look at the summary of the point cloud data we’ll be processing
# processing dirs
c2t_out_dir_temp <- "../data/PFDP_Data/"
psinf_out_dir <- file.path(c2t_out_dir_temp, paste0("point_cloud_processing_delivery_chm",my_chm_res_m,"m") )
las_dir_temp <- "f:\\PFDP_Data\\p4pro_images\\P4Pro_06_17_2021_half_half_optimal\\2_densification\\point_cloud"## class : LAScatalog (v1.2 format 3)
## extent : 499264.2, 499958.8, 4317541, 4318147 (xmin, xmax, ymin, ymax)
## coord. ref. : WGS 84 / UTM zone 13N
## area : 420912.1 m²
## points : 158.01 million points
## type : terrestrial
## density : 375.4 points/m²
## num. files : 1process the point cloud with the cloud2trees::cloud2raster() workflow
# do it
if(!dir.exists(psinf_out_dir)){
# start time
my_st_temp <- Sys.time()
message("starting psinf cloud2raster() step at ...", my_st_temp)
# cloud2trees
psinf_cloud2raster_ans <- cloud2trees::cloud2raster(
output_dir = c2t_out_dir_temp
, input_las_dir = las_dir_temp
, accuracy_level = 2
, keep_intrmdt = T
, dtm_res_m = 0.25
, chm_res_m = my_chm_res_m
, min_height = 0 # effectively generates a DSM based on non-ground points
)
# end
my_end_temp <- Sys.time()
# rename
file.rename(from = file.path(c2t_out_dir_temp,"point_cloud_processing_delivery"), to = psinf_out_dir)
# timer
dplyr::tibble(
timer_cloud2raster_mins = difftime(my_end_temp, my_st_temp, units = c("mins")) %>% as.numeric()
) %>%
readr::write_csv(file = file.path(psinf_out_dir,"processed_tracking_data.csv"), append = F, progress = F)
}else{
dtm_temp <- terra::rast( file.path(psinf_out_dir, "dtm_0.25m.tif") )
chm_temp <- terra::rast( file.path(psinf_out_dir, paste0("chm_", my_chm_res_m,"m.tif")) )
# combine
psinf_cloud2raster_ans <- list(
"dtm_rast" = dtm_temp
, "chm_rast" = chm_temp
)
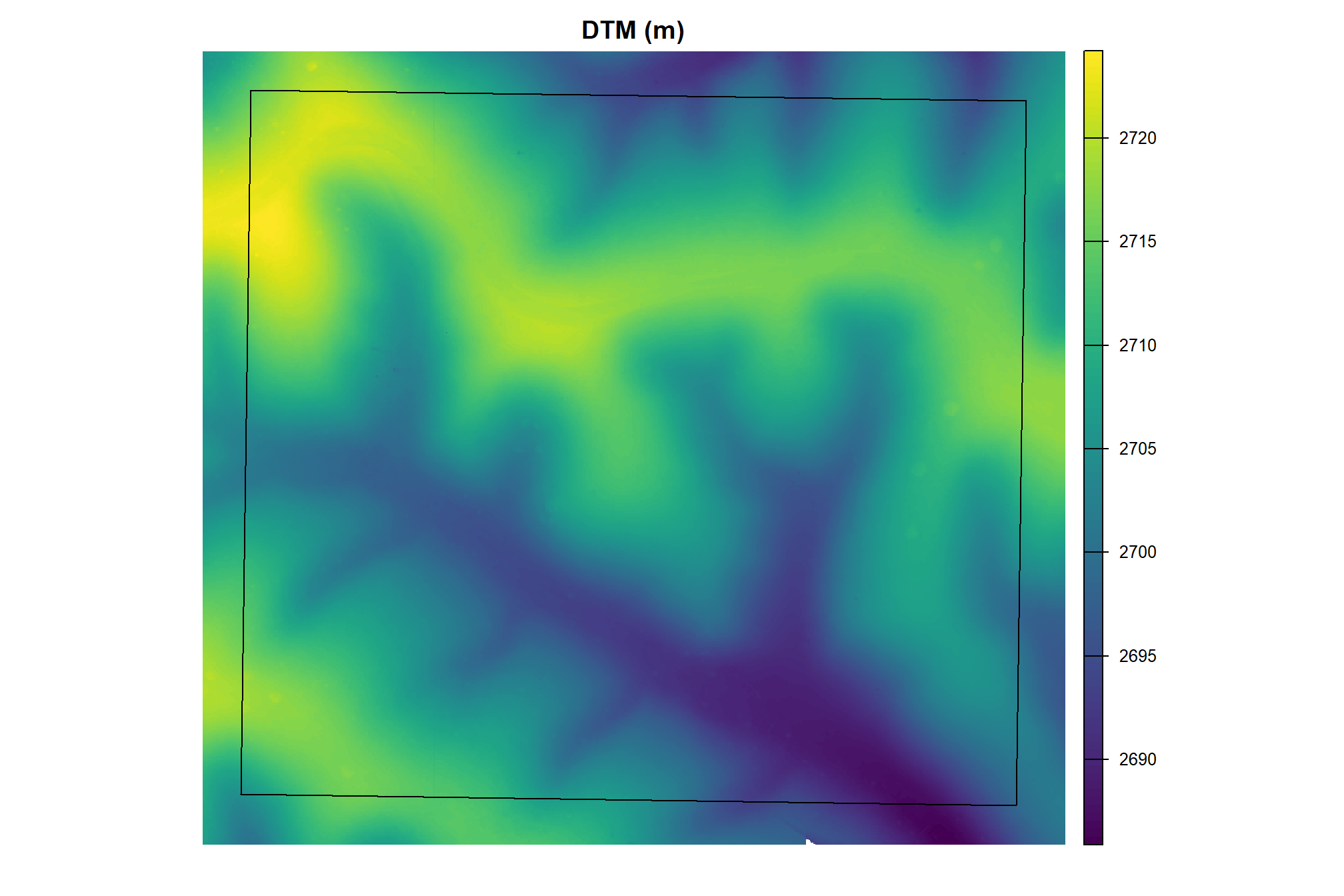
}let’s check out the DTM
psinf_cloud2raster_ans$dtm_rast %>%
terra::crop(
psinf_stand_boundary %>%
sf::st_buffer(22) %>%
sf::st_transform(terra::crs(psinf_cloud2raster_ans$dtm_rast)) %>%
terra::vect()
) %>%
terra::aggregate(fact = 2, na.rm = T, cores = lasR::half_cores()) %>%
terra::plot(
col = viridis::viridis(n = 100)
, axes = F
, main = "DTM (m)"
)
# add stand boundary
terra::plot(
psinf_stand_boundary %>%
sf::st_transform(terra::crs(psinf_cloud2raster_ans$dtm_rast)) %>%
terra::vect()
, add = T, border = "black", col = NA, lwd = 1.2
)
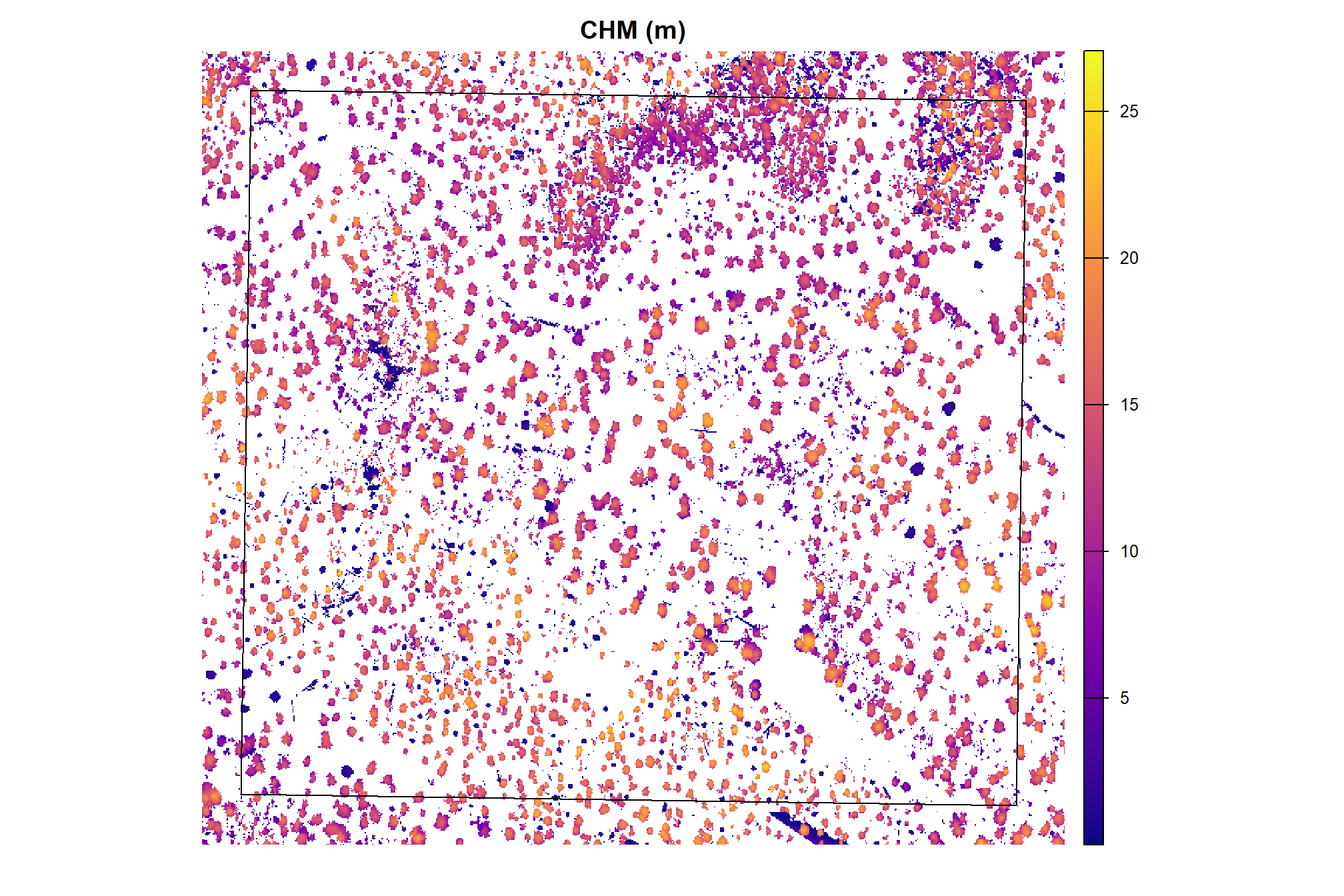
and CHM
psinf_cloud2raster_ans$chm_rast %>%
terra::crop(
psinf_stand_boundary %>%
sf::st_buffer(22) %>%
sf::st_transform(terra::crs(psinf_cloud2raster_ans$chm_rast)) %>%
terra::vect()
) %>%
terra::aggregate(fact = 2, na.rm = T, cores = lasR::half_cores()) %>%
terra::plot(
col = viridis::plasma(100)
, axes = F
, main = "CHM (m)"
)
# add stand boundary
terra::plot(
psinf_stand_boundary %>%
sf::st_transform(terra::crs(psinf_cloud2raster_ans$chm_rast)) %>%
terra::vect()
, add = T, border = "black", col = NA, lwd = 1.2
)
3.1.1.1 DTM and CHM + piles
let’s visually inspect the DTM and CHM with the pile outlines overlaid. we’ll make some functions to use for the different study areas
plt_rast_fn <- function(
rn
, df
, rast
, my_title = ""
, vopt = "viridis"
, lim = NULL
, buff = 10
) {
# scale the buffer based on the largest
d_temp <- df %>%
dplyr::arrange(desc(image_gt_diameter_m)) %>%
dplyr::slice(rn) %>%
sf::st_transform(terra::crs(rast))
# convert rast to df
comp_st <- rast %>%
terra::crop(
d_temp %>% sf::st_buffer(buff) %>% terra::vect()
) %>%
terra::as.data.frame(xy = T) %>%
dplyr::rename(f=3)
# ggplot
comp_temp <- ggplot2::ggplot() +
ggplot2::geom_tile(data = comp_st, ggplot2::aes(x=x,y=y,fill=f)) +
ggplot2::geom_sf(data = d_temp, fill = NA, color = "cyan", lwd = 0.4) +
ggplot2::scale_x_continuous(expand = c(0, 0)) +
ggplot2::scale_y_continuous(expand = c(0, 0)) +
ggplot2::labs(
x = ""
, y = ""
, subtitle = my_title
) +
ggplot2::theme_void() +
ggplot2::theme(
legend.position = "none" # c(0.5,1)
, legend.margin = margin(0,0,0,0)
, legend.text = element_text(size = 8)
, legend.title = element_text(size = 8)
, legend.key = element_rect(fill = "white")
# , plot.title = ggtext::element_markdown(size = 10, hjust = 0.5)
, plot.title = element_text(size = 8, hjust = 0.5, face = "bold")
, plot.subtitle = element_text(size = 6, hjust = 0.5, face = "italic")
)
if(!is.null(lim)){
comp_temp <- comp_temp +
ggplot2::scale_fill_viridis_c(option = vopt, limits = lim)
}else{
comp_temp <- comp_temp +
ggplot2::scale_fill_viridis_c(option = vopt)
}
plt_temp <- comp_temp
return(list("plt"=plt_temp,"d"=d_temp))
}
# plt_rast_fn(rn = 9, df = psinf_slash_piles_polys, rast = psinf_cloud2raster_ans$dtm_rast, vopt = "viridis")
# combine 3
plt_rast_combine <- function(
rn
, df
, dtm_rast
, dtm_rast_vopt = "viridis"
, chm_rast
, chm_rast_vopt = "plasma"
, rgb_rast
, buffer = 10
) {
# composite 1
ans1 <- plt_rast_fn(
rn = rn
, df = df
, rast = dtm_rast
, my_title = "DTM"
, vopt = dtm_rast_vopt
, buff = buffer
)
# composite 2
ans2 <- plt_rast_fn(
rn = rn
, df = df
, rast = chm_rast
, my_title = "CHM"
, vopt = chm_rast_vopt
, buff = buffer
)
# plt rgb
rgb_temp <-
ortho_plt_fn(
rgb_rast = rgb_rast
, stand =ans1$d
, buffer = buffer
, add_stand = F
) +
ggplot2::geom_sf(data = ans1$d, fill = NA, color = "cyan", lwd = 0.3)
# combine
r <- patchwork::wrap_plots(list(rgb_temp,ans1$plt,ans2$plt), nrow = 1)
return(r)
}
# plt_rast_combine(
# rn = 11
# , df = psinf_slash_piles_polys %>% dplyr::filter(is_in_stand)
# , dtm_rast = psinf_cloud2raster_ans$dtm_rast
# , chm_rast = psinf_cloud2raster_ans$chm_rast
# , rgb_rast = psinf_rgb_rast
# )use this handy plotting function for some piles
# add pile locations
plt_list_rast_temp <-
1:sum(psinf_slash_piles_polys$is_in_stand) %>%
sample(
min(10, sum(psinf_slash_piles_polys$is_in_stand))
) %>%
purrr::map(
\(x)
plt_rast_combine(
rn = x
, df = psinf_slash_piles_polys %>% dplyr::filter(is_in_stand)
, dtm_rast = psinf_cloud2raster_ans$dtm_rast
, chm_rast = psinf_cloud2raster_ans$chm_rast
, rgb_rast = psinf_rgb_rast
)
)
# plt_list_rast_temp[[2]]combine the plots with patchwork
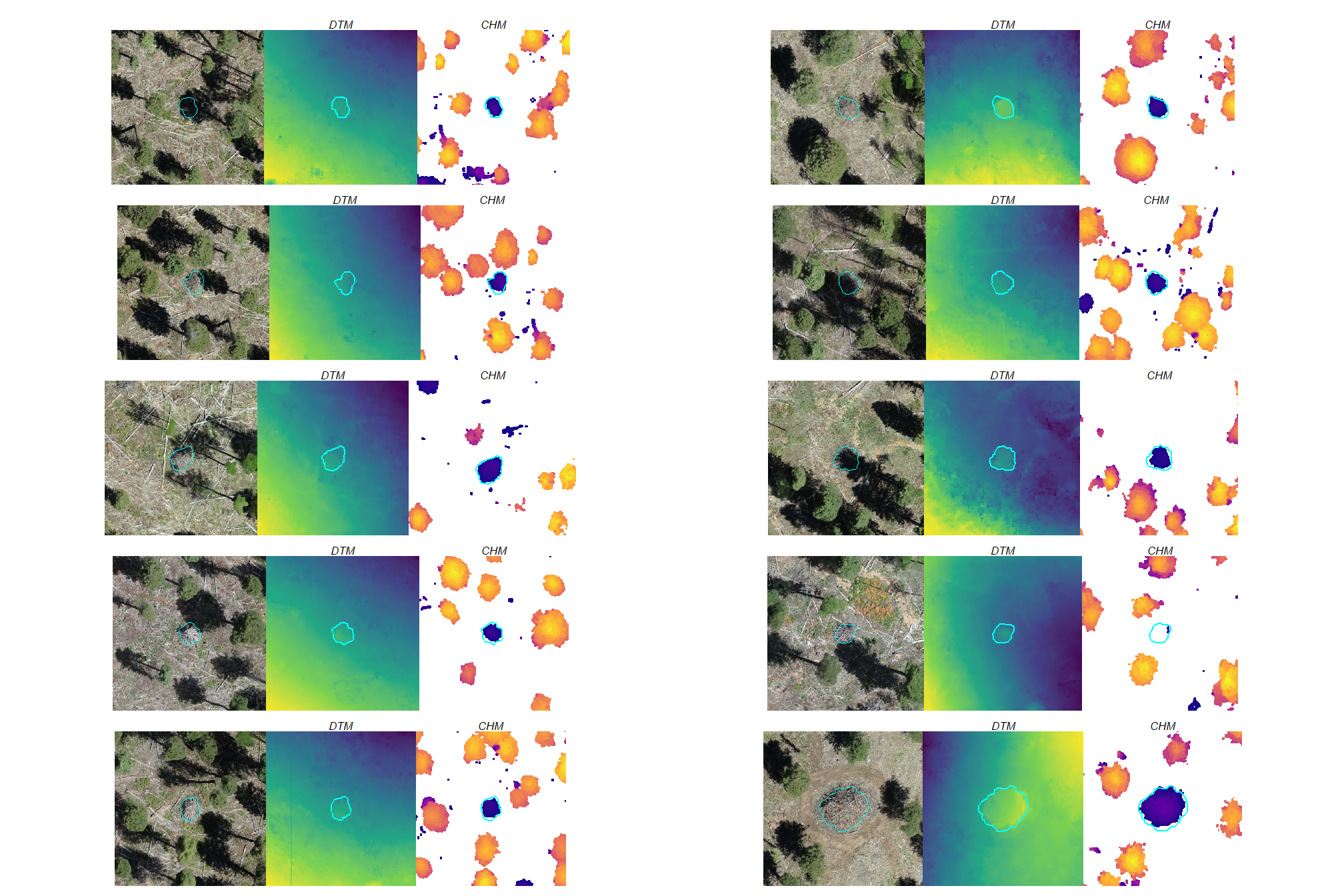
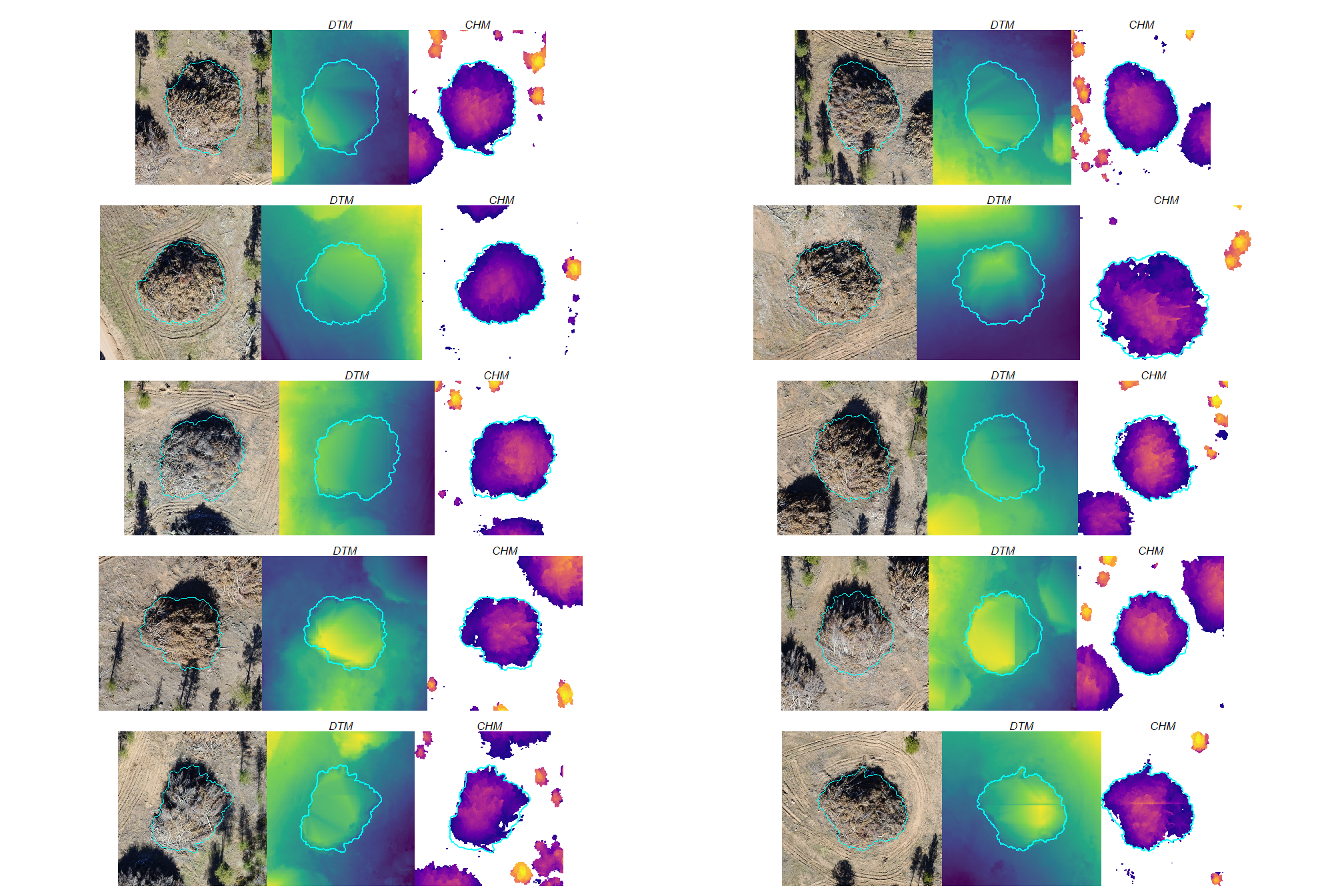
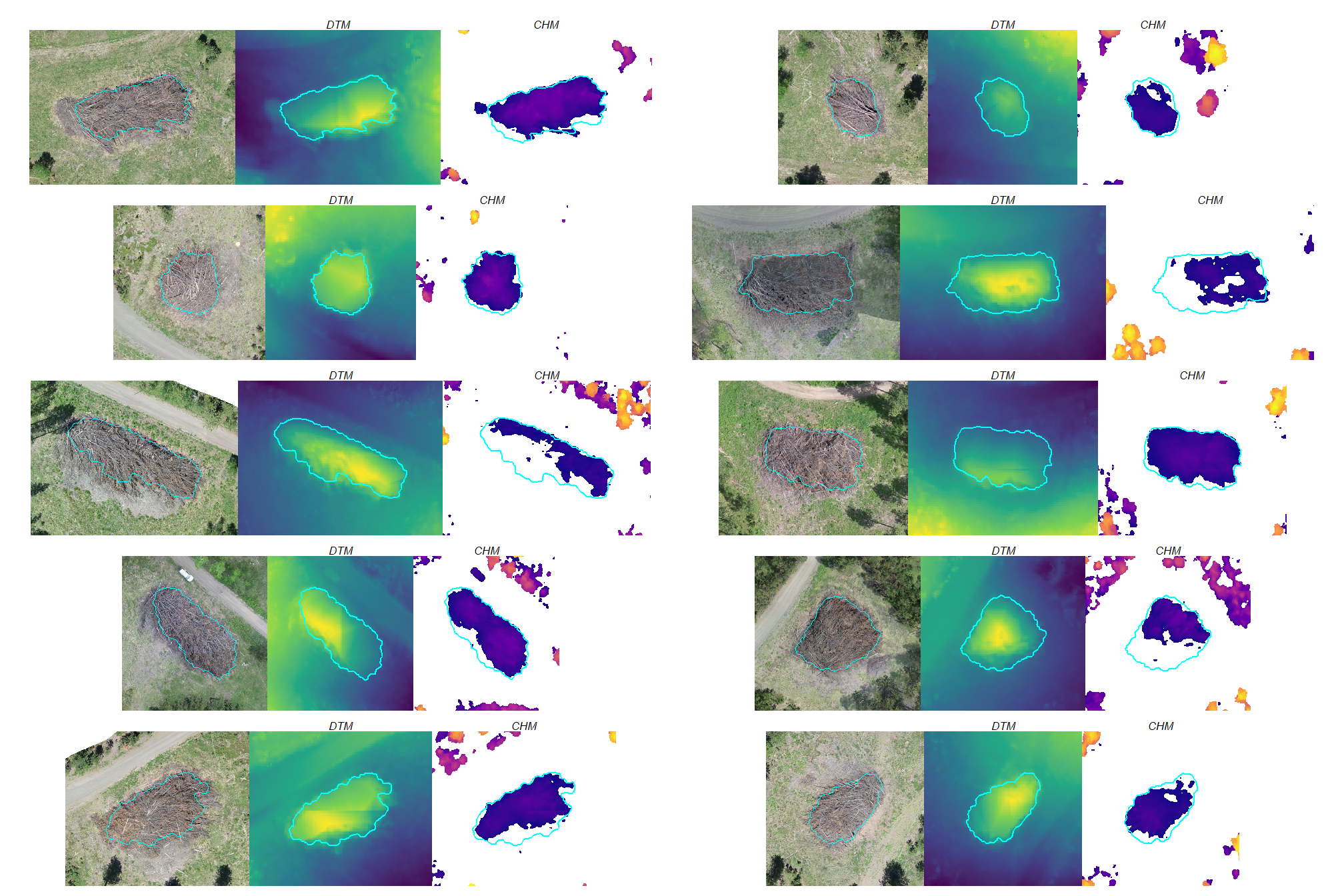
a few things are noteworthy in these examples:
- Piles are clearly visible in some areas of the DTM but less visible in other areas
- Piles are delineated in the CHM but with varying degrees of definition based on surrounding terrain and pile structure
- The DTM and CHM were rarely impacted by shadows in the RGB imagery
- It is interesting to see coarse woody debris occasionally visible in the CHM
3.1.2 TRFO-BLM Pinyon-Juniper Site
for this site, the full point cloud data we have covers a much larger extent than the treatment unit used for this study. we will therefore crop point cloud to a buffered area of interest so that we limit the amount of data processed
###############################################################
# read/crop point cloud
###############################################################
# output dir for clipped las
las_dir_temp <- "../data/Dawson_Data/point_cloud"
if(!dir.exists(las_dir_temp)){dir.create(las_dir_temp, showWarnings = F)}
# do it if not already done
if(dplyr::coalesce(length(list.files(las_dir_temp)),0)<1){
# this reads the metadata from all files in the folder, not the points themselves.
las_ctg_temp <- lidR::readLAScatalog("d:/BLM_CO_SWDF_DawsonFuelsTreatment/Final/PointCloud/Tiles/")
las_ctg_temp
cloud2trees:::check_las_ctg_empty(las_ctg_temp)
# set ctg opts
lidR::opt_select(las_ctg_temp) <- "xyzainrcRGBNC"
lidR::opt_progress(las_ctg_temp) <- T
# write generated results to disk storage rather than keeping everything in memory. This option can be activated with opt_output_files()
lidR::opt_output_files(las_ctg_temp) <- paste0(normalizePath(las_dir_temp),"/", "_{XLEFT}_{YBOTTOM}") # label outputs based on coordinates
lidR::opt_filter(las_ctg_temp) <- "-drop_duplicates"
# clip the point cloud using the polygon and write to a new file
# the lidR::opt_output_files is key to writing the output to disk without loading the entire clipped result into memory.
clip_las_ctg_temp <- lidR::clip_roi(
las = las_ctg_temp
, geometry = pj_stand_boundary %>%
sf::st_buffer(100) %>%
sf::st_bbox() %>%
sf::st_as_sfc() %>%
sf::st_as_sf() %>%
sf::st_transform(sf::st_crs(las_ctg_temp))
)
}else{
clip_las_ctg_temp <- lidR::readLAScatalog(las_dir_temp)
}
# clip_las_ctg_temp
# clip_las_ctg_temp$filenamelook at the summary of the point cloud data we’ll be processing
# processing dirs
c2t_out_dir_temp <- "../data/Dawson_Data/"
pj_out_dir <- file.path(c2t_out_dir_temp, paste0("point_cloud_processing_delivery_chm",my_chm_res_m,"m") )
# las_dir_temp <- las_dir_temp
# read header with catalog
lidR::readLAScatalog(las_dir_temp)## class : LAScatalog (v1.2 format 2)
## extent : 707813.6, 708409.9, 4193155, 4193638 (xmin, xmax, ymin, ymax)
## coord. ref. : NAD83(2011) / UTM zone 12N
## area : 288106.3 m²
## points : 163.84 million points
## type : terrestrial
## density : 568.7 points/m²
## density : 568.7 pulses/m²
## num. files : 1process the point cloud with the cloud2trees::cloud2raster() workflow
# do it
if(!dir.exists(pj_out_dir)){
# start time
my_st_temp <- Sys.time()
message("starting pj cloud2raster() step at ...", my_st_temp)
# cloud2trees
pj_cloud2raster_ans <- cloud2trees::cloud2raster(
output_dir = c2t_out_dir_temp
, input_las_dir = las_dir_temp
, accuracy_level = 2
, keep_intrmdt = T
, dtm_res_m = 0.2
, chm_res_m = my_chm_res_m
, min_height = 0 # effectively generates a DSM based on non-ground points
)
# end
my_end_temp <- Sys.time()
# rename
file.rename(from = file.path(c2t_out_dir_temp,"point_cloud_processing_delivery"), to = pj_out_dir)
# timer
dplyr::tibble(
timer_cloud2raster_mins = difftime(my_end_temp, my_st_temp, units = c("mins")) %>% as.numeric()
) %>%
readr::write_csv(file = file.path(pj_out_dir,"processed_tracking_data.csv"), append = F, progress = F)
}else{
dtm_temp <- terra::rast( file.path(pj_out_dir, "dtm_0.2m.tif") ) %>%
terra::aggregate(fact = 2, na.rm=T, cores = lasR::half_cores())
chm_temp <- terra::rast( file.path(pj_out_dir, paste0("chm_", my_chm_res_m,"m.tif")) )
# combine
pj_cloud2raster_ans <- list(
"dtm_rast" = dtm_temp
, "chm_rast" = chm_temp
)
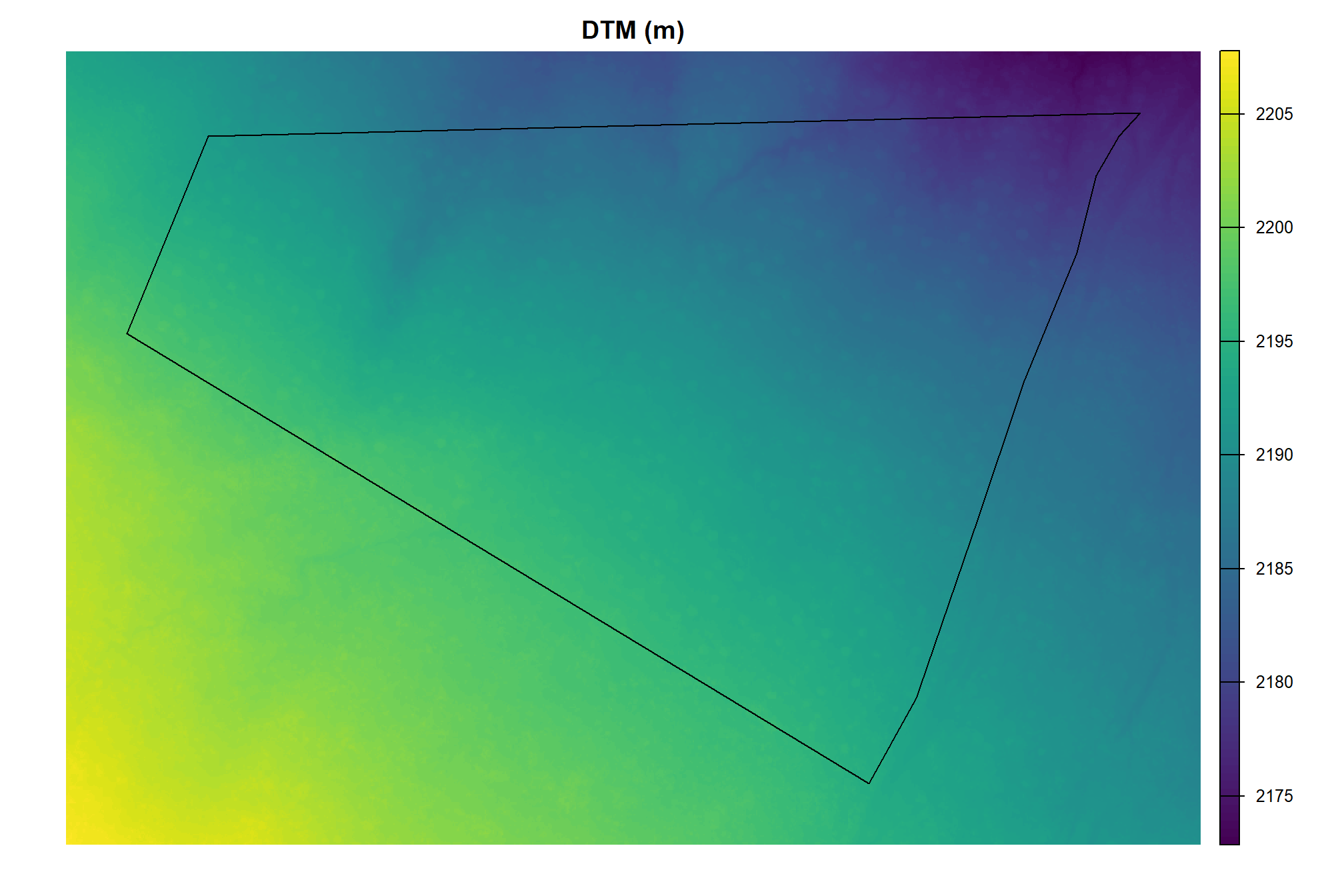
}let’s check out the DTM
pj_cloud2raster_ans$dtm_rast %>%
terra::crop(
pj_stand_boundary %>%
sf::st_buffer(22) %>%
sf::st_transform(terra::crs(pj_cloud2raster_ans$dtm_rast)) %>%
terra::vect()
) %>%
terra::plot(
col = viridis::viridis(n = 100)
, axes = F
, main = "DTM (m)"
)
# add stand boundary
terra::plot(
pj_stand_boundary %>%
sf::st_transform(terra::crs(pj_cloud2raster_ans$dtm_rast)) %>%
terra::vect()
, add = T, border = "black", col = NA, lwd = 1.2
)
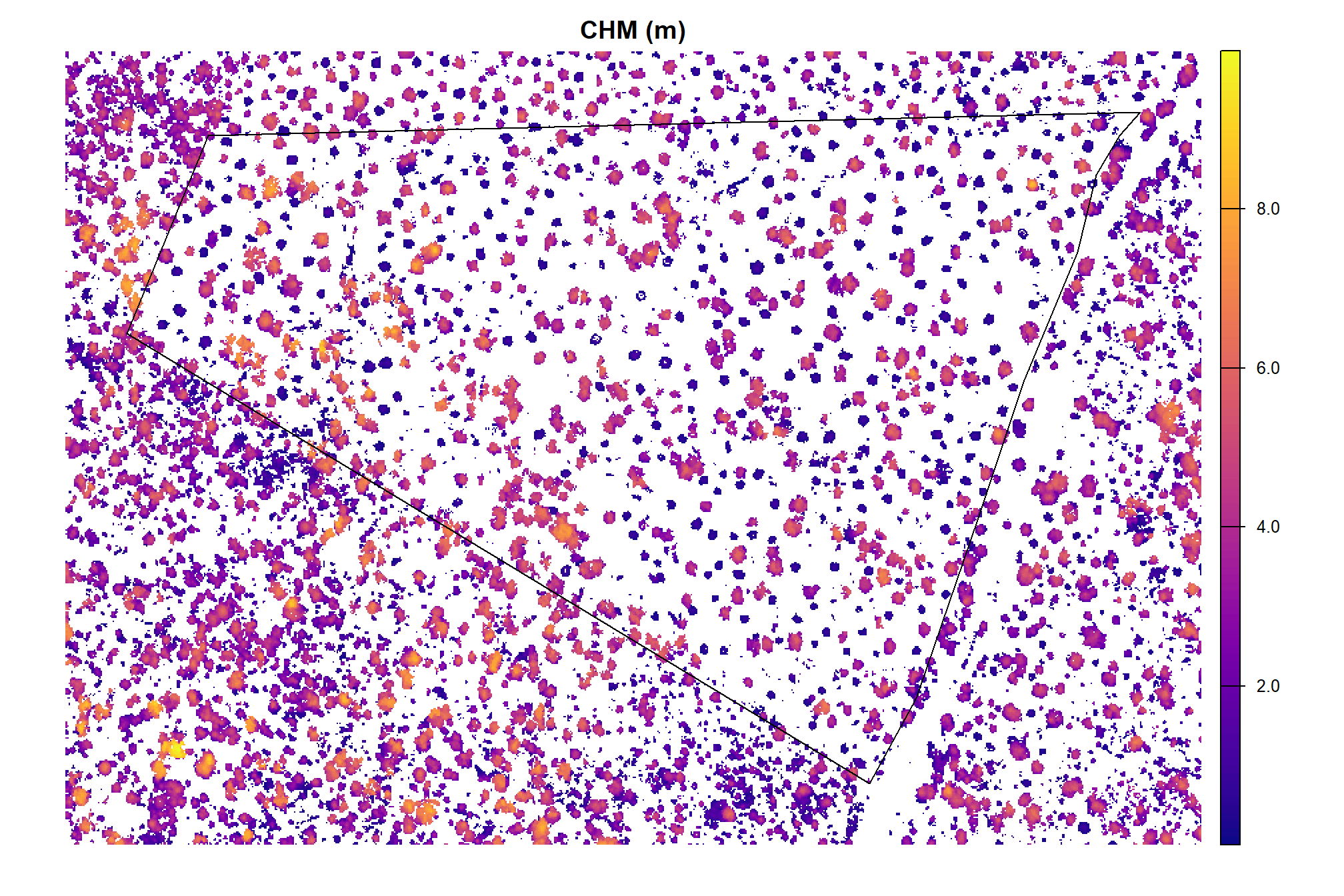
and CHM
pj_cloud2raster_ans$chm_rast %>%
terra::crop(
pj_stand_boundary %>%
sf::st_buffer(22) %>%
sf::st_transform(terra::crs(pj_cloud2raster_ans$chm_rast)) %>%
terra::vect()
) %>%
terra::aggregate(fact = 2, na.rm = T, cores = lasR::half_cores()) %>%
terra::plot(
col = viridis::plasma(100)
, axes = F
, main = "CHM (m)"
)
# add stand boundary
terra::plot(
pj_stand_boundary %>%
sf::st_transform(terra::crs(pj_cloud2raster_ans$chm_rast)) %>%
terra::vect()
, add = T, border = "black", col = NA, lwd = 1.2
)
3.1.2.1 DTM and CHM + piles
let’s visually inspect the DTM and CHM with the pile outlines overlaid for a sample of piles
# add pile locations
plt_list_rast_temp <-
1:sum(pj_slash_piles_polys$is_in_stand) %>%
sample(
min(10, sum(pj_slash_piles_polys$is_in_stand))
) %>%
purrr::map(
\(x)
plt_rast_combine(
rn = x
, df = pj_slash_piles_polys %>% dplyr::filter(is_in_stand)
, dtm_rast = pj_cloud2raster_ans$dtm_rast
, chm_rast = pj_cloud2raster_ans$chm_rast
, rgb_rast = pj_rgb_rast
)
)
# plt_list_rast_temp[[2]]combine the plots with patchwork
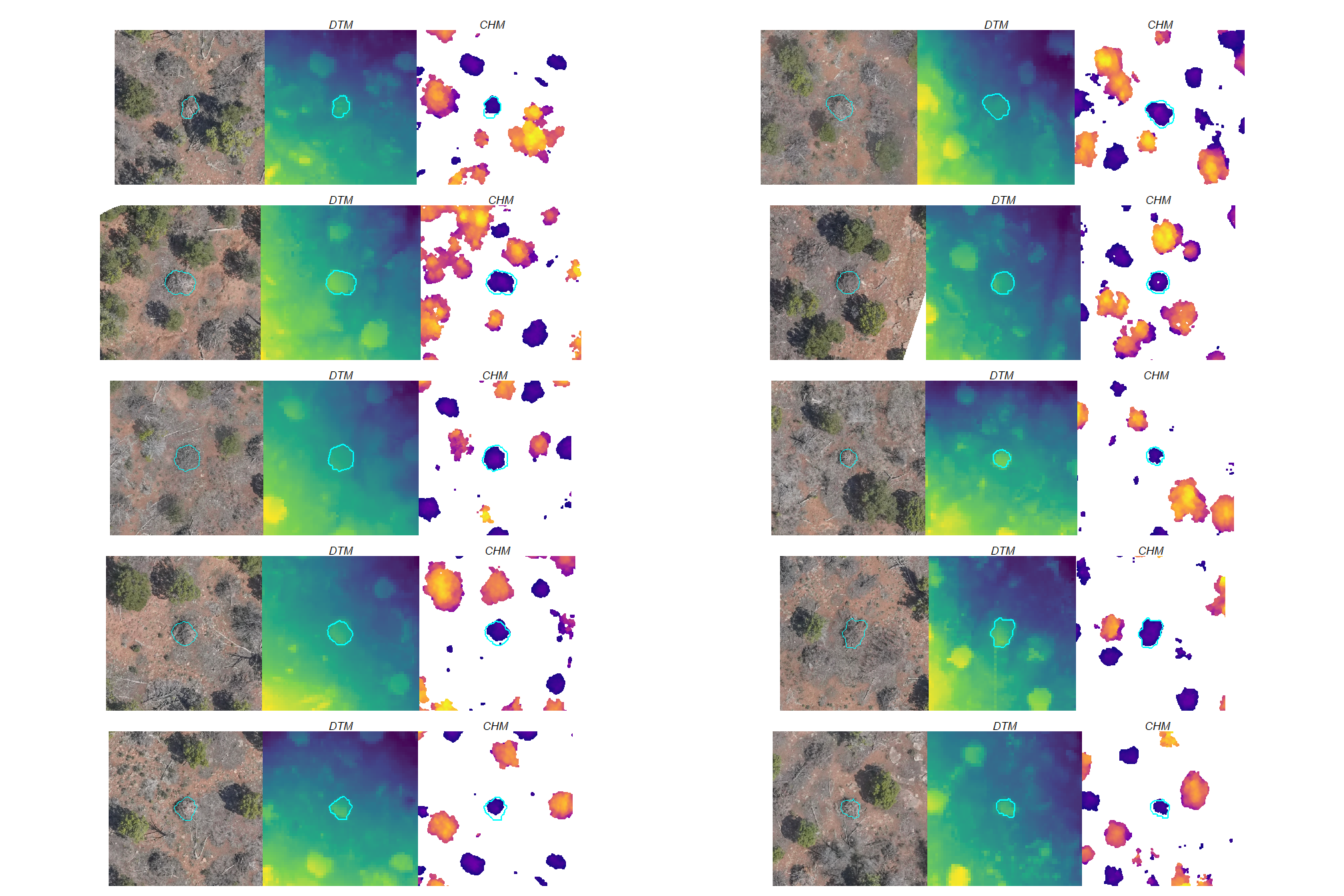
3.1.3 BHEF Ponderosa Pine Site
for this site, the point cloud files are dispersed across different folders. so, we’ll manually specify which files to use.
look at the summary of the point cloud data we’ll be processing
# processing dirs
c2t_out_dir_temp <- "../data/BHEF_202306/"
bhef_out_dir <- file.path(c2t_out_dir_temp, paste0("point_cloud_processing_delivery_chm",my_chm_res_m,"m") )
# las_dir_temp <- "where/are/the/pointcloud"
las_flist_temp <- c(
"F:\\UAS_Collections\\BHEF_202306/Unit1and3Processing.files/BHEF_202306_Unit1and3_laz.laz"
, "F:\\UAS_Collections\\BHEF_202306/Unit2Processing.files/BHEF_202306_Unit2_laz.laz"
, list.files(
"F:\\UAS_Collections\\BHEF_202306\\00good_simplified_point_clouds_reduced_overlap"
, pattern = ".*\\.(laz|las)$"
, full.names = TRUE, recursive = T
)
)## class : LAScatalog (v1.2 format 2)
## extent : 608234, 610968.2, 4888216, 4889418 (xmin, xmax, ymin, ymax)
## coord. ref. : NAD83 / UTM zone 13N
## area : 3.48 km²
## points : 1.42 billion points
## type : airborne
## density : 408 points/m²
## density : 408 pulses/m²
## num. files : 7process the point cloud with the cloud2trees::cloud2raster() workflow
# do it
if(!dir.exists(bhef_out_dir)){
# start time
my_st_temp <- Sys.time()
message("starting bhef cloud2raster() step at ...", my_st_temp)
# cloud2trees
bhef_cloud2raster_ans <- cloud2trees::cloud2raster(
output_dir = c2t_out_dir_temp
, input_las_dir = las_flist_temp
, accuracy_level = 2
, keep_intrmdt = T
, dtm_res_m = 0.2
, chm_res_m = my_chm_res_m
, min_height = 0 # effectively generates a DSM based on non-ground points
)
# end
my_end_temp <- Sys.time()
# rename
file.rename(from = file.path(c2t_out_dir_temp,"point_cloud_processing_delivery"), to = bhef_out_dir)
# timer
dplyr::tibble(
timer_cloud2raster_mins = difftime(my_end_temp, my_st_temp, units = c("mins")) %>% as.numeric()
) %>%
readr::write_csv(file = file.path(bhef_out_dir,"processed_tracking_data.csv"), append = F, progress = F)
}else{
dtm_temp <- terra::rast( file.path(bhef_out_dir, "dtm_0.2m.tif") ) %>%
terra::aggregate(fact = 2, na.rm=T, cores = lasR::half_cores())
chm_temp <- terra::rast( file.path(bhef_out_dir, paste0("chm_", my_chm_res_m,"m.tif")) )
# combine
bhef_cloud2raster_ans <- list(
"dtm_rast" = dtm_temp
, "chm_rast" = chm_temp
)
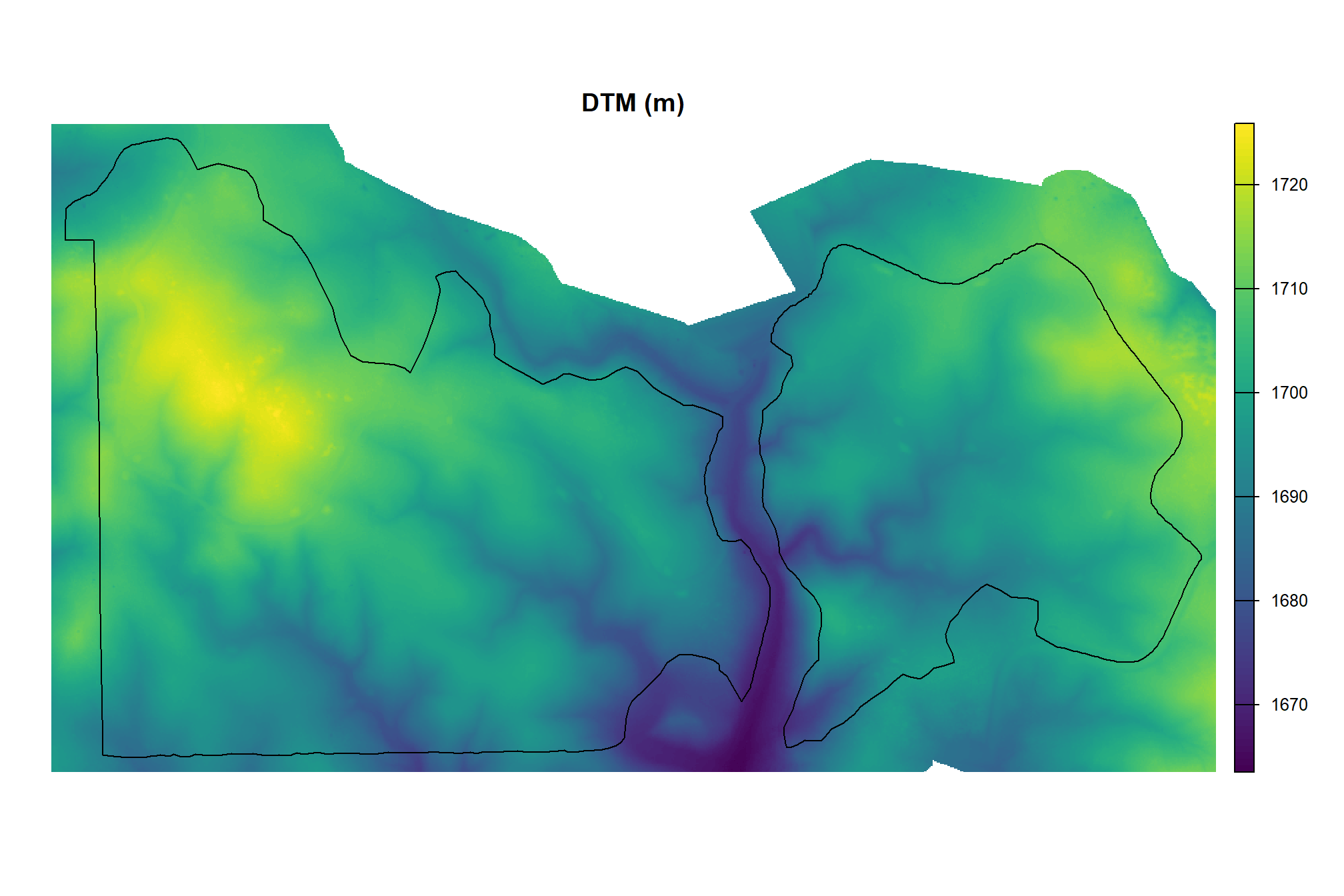
}let’s check out the DTM
bhef_cloud2raster_ans$dtm_rast %>%
terra::crop(
bhef_stand_boundary %>%
sf::st_buffer(22) %>%
sf::st_transform(terra::crs(bhef_cloud2raster_ans$dtm_rast)) %>%
terra::vect()
) %>%
terra::plot(
col = viridis::viridis(n = 100)
, axes = F
, main = "DTM (m)"
)
# add stand boundary
terra::plot(
bhef_stand_boundary %>%
sf::st_transform(terra::crs(bhef_cloud2raster_ans$dtm_rast)) %>%
terra::vect()
, add = T, border = "black", col = NA, lwd = 1.2
)
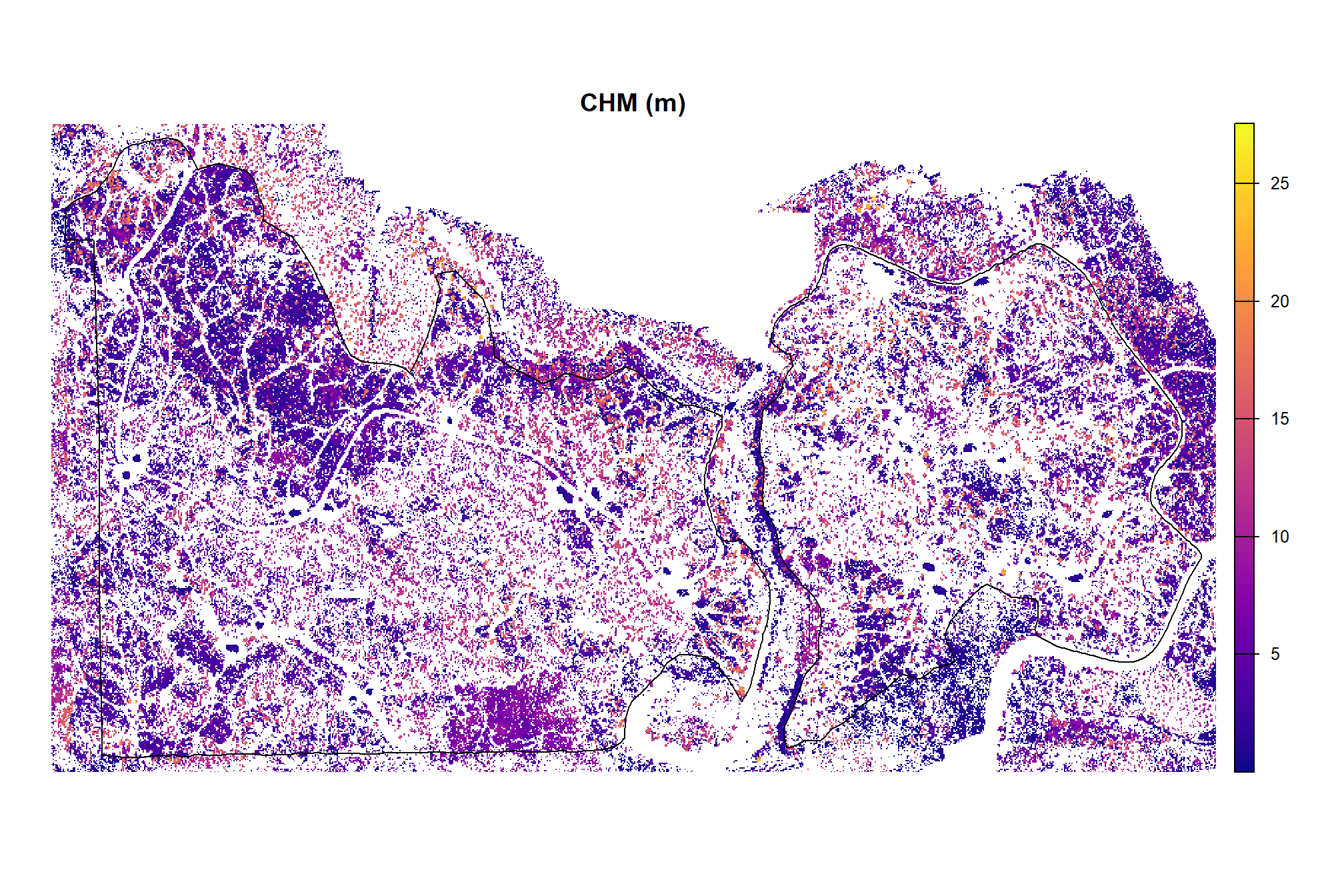
and CHM
bhef_cloud2raster_ans$chm_rast %>%
terra::crop(
bhef_stand_boundary %>%
sf::st_buffer(22) %>%
sf::st_transform(terra::crs(bhef_cloud2raster_ans$chm_rast)) %>%
terra::vect()
) %>%
terra::aggregate(fact = 2, na.rm = T, cores = lasR::half_cores()) %>%
terra::plot(
col = viridis::plasma(100)
, axes = F
, main = "CHM (m)"
)
# add stand boundary
terra::plot(
bhef_stand_boundary %>%
sf::st_transform(terra::crs(bhef_cloud2raster_ans$chm_rast)) %>%
terra::vect()
, add = T, border = "black", col = NA, lwd = 1.2
)
3.1.3.1 DTM and CHM + piles
let’s visually inspect the DTM and CHM with the pile outlines overlaid for a sample of piles
# add pile locations
plt_list_rast_temp <-
1:sum(bhef_slash_piles_polys$is_in_stand) %>%
sample(
min(10, sum(bhef_slash_piles_polys$is_in_stand))
) %>%
purrr::map(
\(x)
plt_rast_combine(
rn = x
, df = bhef_slash_piles_polys %>% dplyr::filter(is_in_stand)
, dtm_rast = bhef_cloud2raster_ans$dtm_rast
, chm_rast = bhef_cloud2raster_ans$chm_rast
, rgb_rast = bhef_rgb_rast
)
)
# plt_list_rast_temp[[2]]combine the plots with patchwork
3.1.4 ARNF Ponderosa Pine Site
look at the summary of the point cloud data we’ll be processing
# processing dirs
c2t_out_dir_temp <- "../data/ARNF_DiamondView_202510/" # where do you want to save processed data to?
arnf_out_dir <- file.path(c2t_out_dir_temp, paste0("point_cloud_processing_delivery_chm",my_chm_res_m,"m") )
las_dir_temp <- "F:/UAS_Collections/ARNF_DiamondView_202510/point_cloud_high_density" # where is the raw las## class : LAScatalog (v1.2 format 3)
## extent : 455073.8, 456582.3, 4535127, 4536067 (xmin, xmax, ymin, ymax)
## coord. ref. : WGS 84 / UTM zone 13N
## area : 1.24 km²
## points : 1.37 billion points
## type : airborne
## density : 1102.8 points/m²
## density : 39.9 pulses/m²
## num. files : 44process the point cloud with the cloud2trees::cloud2raster() workflow
# do it
if(!dir.exists(arnf_out_dir)){
# start time
my_st_temp <- Sys.time()
message("starting arnf cloud2raster() step at ...", my_st_temp)
# cloud2trees
arnf_cloud2raster_ans <- cloud2trees::cloud2raster(
output_dir = c2t_out_dir_temp
, input_las_dir = las_dir_temp
, accuracy_level = 2
, keep_intrmdt = T
, dtm_res_m = 0.2
, chm_res_m = my_chm_res_m
, min_height = 0 # effectively generates a DSM based on non-ground points
)
# end
my_end_temp <- Sys.time()
# rename
file.rename(from = file.path(c2t_out_dir_temp,"point_cloud_processing_delivery"), to = arnf_out_dir)
# timer
dplyr::tibble(
timer_cloud2raster_mins = difftime(my_end_temp, my_st_temp, units = c("mins")) %>% as.numeric()
) %>%
readr::write_csv(file = file.path(arnf_out_dir,"processed_tracking_data.csv"), append = F, progress = F)
}else{
dtm_temp <- terra::rast( file.path(arnf_out_dir, "dtm_0.2m.tif") ) %>%
terra::aggregate(fact = 2, na.rm=T, cores = lasR::half_cores())
chm_temp <- terra::rast( file.path(arnf_out_dir, paste0("chm_", my_chm_res_m,"m.tif")) )
# combine
arnf_cloud2raster_ans <- list(
"dtm_rast" = dtm_temp
, "chm_rast" = chm_temp
)
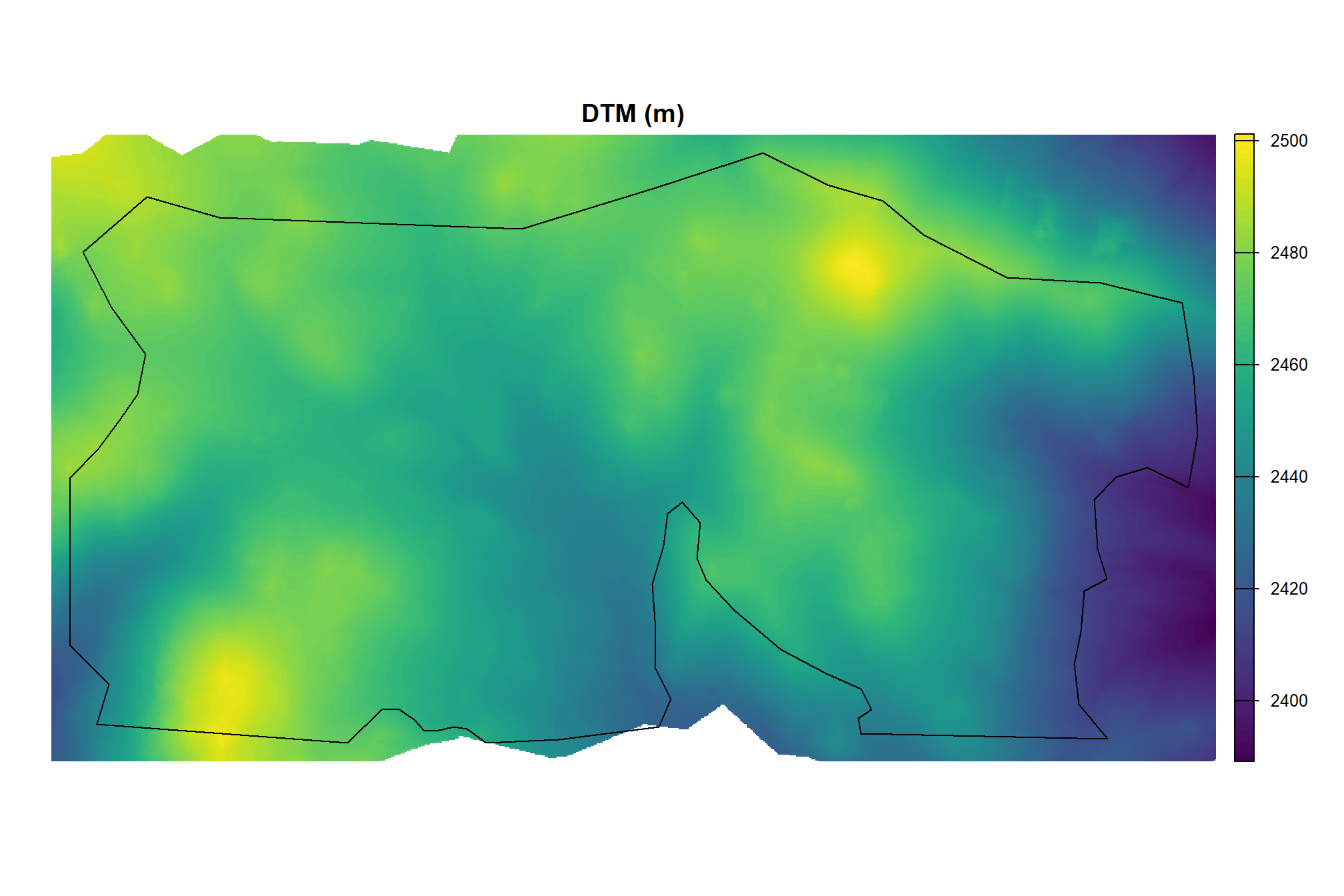
}let’s check out the DTM
arnf_cloud2raster_ans$dtm_rast %>%
terra::crop(
arnf_stand_boundary %>%
sf::st_buffer(22) %>%
sf::st_transform(terra::crs(arnf_cloud2raster_ans$dtm_rast)) %>%
terra::vect()
) %>%
terra::plot(
col = viridis::viridis(n = 100)
, axes = F
, main = "DTM (m)"
)
# add stand boundary
terra::plot(
arnf_stand_boundary %>%
sf::st_transform(terra::crs(arnf_cloud2raster_ans$dtm_rast)) %>%
terra::vect()
, add = T, border = "black", col = NA, lwd = 1.2
)
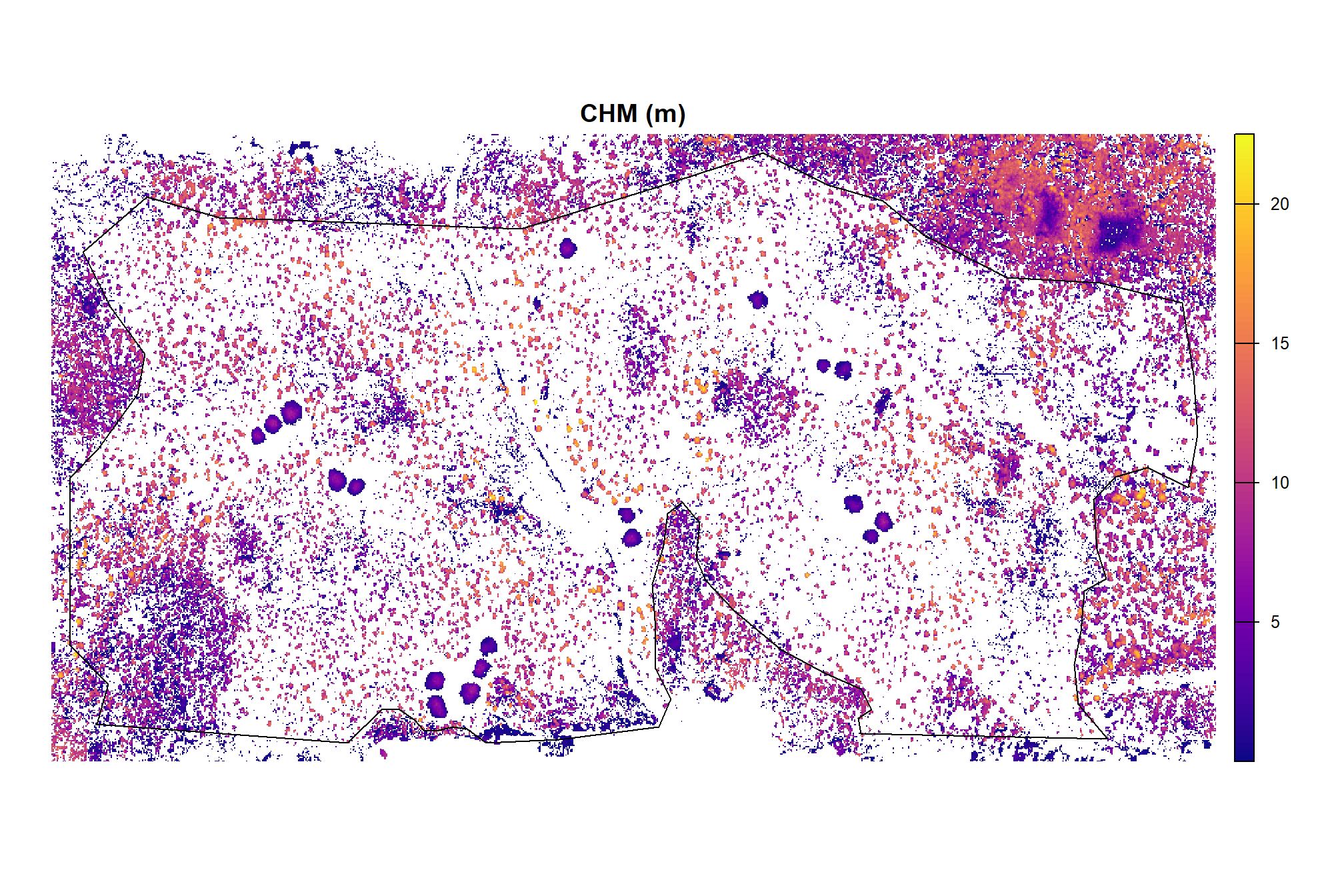
and CHM
arnf_cloud2raster_ans$chm_rast %>%
terra::crop(
arnf_stand_boundary %>%
sf::st_buffer(22) %>%
sf::st_transform(terra::crs(arnf_cloud2raster_ans$chm_rast)) %>%
terra::vect()
) %>%
terra::aggregate(fact = 2, na.rm = T, cores = lasR::half_cores()) %>%
terra::plot(
col = viridis::plasma(100)
, axes = F
, main = "CHM (m)"
)
# add stand boundary
terra::plot(
arnf_stand_boundary %>%
sf::st_transform(terra::crs(arnf_cloud2raster_ans$chm_rast)) %>%
terra::vect()
, add = T, border = "black", col = NA, lwd = 1.2
)
3.1.4.1 DTM and CHM + piles
let’s visually inspect the DTM and CHM with the pile outlines overlaid for a sample of piles
# add pile locations
plt_list_rast_temp <-
1:sum(arnf_slash_piles_polys$is_in_stand) %>%
sample(
min(10, sum(arnf_slash_piles_polys$is_in_stand))
) %>%
purrr::map(
\(x)
plt_rast_combine(
rn = x
, df = arnf_slash_piles_polys %>% dplyr::filter(is_in_stand)
, dtm_rast = arnf_cloud2raster_ans$dtm_rast
, chm_rast = arnf_cloud2raster_ans$chm_rast
, rgb_rast = arnf_rgb_rast
)
)
# plt_list_rast_temp[[2]]combine the plots with patchwork